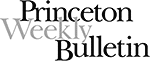Princeton University
Princeton Weekly Bulletin February 5, 2007, Vol. 96, No. 14 prev next current
- Page One
- • Trustees hold the line on tuition, approve funding for key initiatives
- • Scientists build a world in a grain of silicon
- Inside
- • Wilson School expands Scholars in the Nation’s Service Initiative
- • Celebrating music’s inspirational power
- • A window onto Whitman College
- People
- • Martin Kruskal, pre-eminent mathematician, dies at age 81
- • Former Congressman Leach joins Wilson School faculty
- • Spotlight, briefs
- Almanac
- • Calendar of events
- • Nassau notes
- • By the numbers
- The Bulletin is published weekly during the academic year, except during University breaks and exam weeks, by the Office of Communications. Second class postage paid at Princeton. Postmaster: Send address changes to Princeton Weekly Bulletin, Office of Communications, Princeton University, 22 Chambers St., Suite 201, Princeton, NJ 08542. Permission is given to adapt, reprint or excerpt material from the Bulletin for use in other media.
- Subscriptions. The Bulletin is distributed free to faculty, staff and students. Others may subscribe to the Bulletin for $30 for the 2006-07 academic year (half price for current Princeton parents and people over 65). Send a check to Office of Communications, Princeton University, 22 Chambers St., Suite 201, Princeton, NJ 08542.
- Deadlines. In general, the copy deadline for each issue is the Friday 10 days in advance of the Monday cover date. The deadline for the Bulletin that covers Feb. 19-25 is Friday, Feb. 9. A complete publication schedule is available at www.princeton.edu/ pr/ pwb/ deadlines.html; or by calling (609) 258-3601.
- Editor: Ruth Stevens Calendar editor: Shani Hilton Staff writers: Jennifer Greenstein Altmann, Eric Quiñones Contributing writers: Chad Boutin Photographers: Denise Applewhite, John Jameson Design: Maggie Westergaard Web edition: Mahlon Lovett
Scientists build a world in a grain of silicon

This artist’s conception shows a portion of the tiny ecosystem the team created for a population of E. coli bacteria. Each square chamber has 100 micrometer sides and is connected to its neighbors by a tunnel half that long. Narrow tubes, represented by the lines above and below each chamber, carry nutrients in and waste materials out. By varying the nutrient flow into each chamber, the scientists can mimic a real-world habitat with different resources in different areas, permitting the study of ecology and adaptation in a laboratory setting. (rendering: courtesy of the Austin Group)
Princeton NJ — Ever since Charles Darwin proposed that animals adapt to their environment, scientists have dreamed of experimenting with this theory in a real-world landscape. Holding them back was the difficulty of creating a complex ecosystem that could be manipulated and controlled without placing wildlife at risk.
Now, Princeton scientists have found a way around this problem by fashioning a living, changeable ecosystem out of a tiny chip of silicon. Their creation is one of the strangest and smallest environments ever seen, but it could provide a valuable model to help researchers better understand how organisms survive in the natural world.
The spartan landscape is a single row of 85 microscopic chambers connected by narrow corridors, but it possesses most of the complexities of an environment many times its size. In this miniature habitat, the organisms compete for resources, strike out for new territories and adapt to the different resources available. It all happens just as it might among, say, a population of giant pandas. But because the only creatures that live here are common E. coli bacteria, scientists can watch and even manipulate these events without risk to any endangered species.
“These bacteria adapt, and perhaps evolve, as they learn to live in the complex world we have made,” said Robert Austin, professor of physics and senior colleague in the research effort. “Over time, they restrict their growth in places with lots of food to avoid overpopulation, and they learn to grow in places that are poor in resources but have lots of space. There is an analogy to how ‘higher’ organisms such as humans might expand on the surface of the Earth and adapt to their local environment.”
Experiments with the artificial landscape have already netted the team publication of a paper in the Nov. 7, 2006, issue of the Proceedings of the National Academy of Sciences. The paper demonstrates that the team’s creation offers a window into the world of adaptation, but the researchers’ results have a deeper significance. The dynamics of the habitat could permit the scientists to develop bacteria with specific talents (see sidebar). The work also shows that a cross-disciplinary approach to science can sometimes lead to a long-sought solution to a problem.
To breed a champion
While the tiny habitat created by Robert Austin’s group already has revealed insights into the fundamentals of adaptive ecology, it could someday offer practical applications as well.
The team is exploring how it can be used to breed useful strains of bacteria, much as one might breed dogs for hunting or shepherding.
For example, some bacteria give off pure hydrogen gas as a waste product, and thus could prove valuable as a safe, sustainable source of fuel. A strain of bacteria that produces high volumes of the gas could bring scientists a step closer to the much-touted hydrogen economy. The team’s adaptive landscape may enable researchers to develop a strain with such characteristics.
“The basic idea is directed evolution,” Austin said. “By observing the growth of different groups of bacteria in different chambers, we can also monitor each chamber for a desirable product, in this case hydrogen gas. We can reward those populations that produce lots of gas by giving them more food and space. Conversely, we would ‘punish’ underachieving bacterial colonies, but would not destroy them.”
The limited contact among the various colonies would allow strains with different talent at hydrogen production to emerge, and the occasional cross-breeding between natives and plucky colonists could bring beneficial traits into the colony’s overall genetic makeup. Eventually, a highly productive strain would emerge.
“In this way we can direct the evolution of bacteria in the way we want,” Austin said. “We can let fitness selection guide the evolution of a species toward our externally determined goal.”
This part of the team’s effort is a close collaboration with Charles Dismukes’ group in the Department of Chemistry as part of the national BioSolarH2 project that Dismukes leads.
Austin added that their findings also might have application in the realm of nanotechnology, the same methodology that enabled the team to create the habitat in the first place.
“We might even use these techniques to ‘evolve’ new substances as well as new kinds of bacteria,” he said. “Nanotechnologists are often interested in molecules that self-assemble, and we might be able to direct their assembly by observing the steps by which they come together and change what we like. It’s just pie in the sky right now, but we do have funding from the Defense Advanced Research Projects Agency to pursue it.”
“We are all nominally physicists in this group, but creating the habitat has in essence allowed us all to become experimental ecologists,” said Juan Keymer, lead author of the paper and one of the postdoctoral researchers in Austin’s group, which specializes in biological physics. “Historically, a few physicists have crossed over into ecology, but it’s all been theoretical work. Now we can bridge the gap into the biological world.”
Physicists are taught to work with particles that interact according to rules, as they do in a nuclear reactor or a particle accelerator, Keymer said, allowing them to model particle behavior with computers. Though the physicists have adapted their computer models to consider predator-prey interactions in a similar light, the models can only go so far describing relationships among complex living organisms.
“This is the first time we have been able to work with a real ecosystem and living creatures,” said Peter Galajda, another postdoctoral researcher. “It’s both literally and figuratively a whole new world for us.”
This new world is constructed of silicon — the same material used for computer chips — and the team’s experience with nanotechnology and microfabrication techniques made it possible. Each of the 85 “neighborhoods” is a square chamber 100 micrometers to a side and 30 micrometers deep, providing enough elbow room for about 10,000 E. coli at most. Food and waste are introduced and removed through channels too narrow for the bacteria to pass through, and by varying the channels’ flow rates, the team can make life easier or harder on a chamber’s residents.
Should a bacterium find life in one chamber inhospitable, escape routes beckon: Two narrow corridors lead through opposite walls into the adjacent chambers. But the corridors are 50 micrometers long — a decent hike by bacterial standards — and any would-be colonists might have to traverse several relatively poor areas before finding one that seems livable. While the new residents try to make the best of their surroundings, their offspring adapt even more. They often thrive in a chamber that would have been marginally comfortable for their ancestors. Ecologists would say they have adapted to a different “niche” in the landscape.
“These sorts of varied ecological ‘niches’ exist for all species everywhere in nature, from the birds in the Galápagos Islands to the humans in the heart of New York City,” Keymer said. “Adaptation is of course thought to be the first step in the development of new species, though watching it happen has proved elusive. Our artificial landscape may provide the opportunity to observe these changes, though, because of the way it allows the bacteria to colonize different areas over many generations.”
When several farflung populations of a species adapt to different niche environments, all of which have very limited contact with one another because of the difficulty of traveling between them, the species forms what ecologists call a “metapopulation” — a group of groups. Such a split has been observed in large animal populations, such as among the finches Darwin observed on different islands in the Galápagos. But never before this experiment had a metapopulation been seen among bacteria.
“It is this limited contact among groups in different niches that seems to create an environment that encourages evolution to occur,” Keymer said. “Two widely divergent environments will breed two populations suited to each — that’s well understood. But if the two environments are opposites, and there’s a street connecting them, the cells traveling between them will constantly frustrate the process. The evolution will be faster and more dynamic.”
The group is not sure yet whether their habitat has actually produced evolutionary change among the E. coli, or whether the bacteria have simply adapted well to different niches. Finding ways to investigate this question is one of their next goals. In the meantime, Keymer said, the tiny world has proven its worth as a testing ground.
“We have already been able to demonstrate that the ecology of bacteria has a great deal of parallels with that of larger animals,” he said. “We are seeing behaviors that are common to creatures large and small. That’s encouraging, as it means we’ve already got a good model of the natural world that we can work to improve.”